Title: Regulatory paradigm of Dscam1 stochastic alternative splicing through conserved long-range RNA structures
Haiyang Dong, Bingbing Xu, Lili Wu, Zhechao Wang, Zedong Li, Jinpeng Xie, Jingsong Lv, Wanqing Yang, Nengcheng Bao, Huiru Qu, Jiayan Fu, Ru Yan, Yadong Wang, Lei Li, Feng Shi, Jianhua Huang, Xiaofeng Yang, Yongfeng Jin*
Abstract
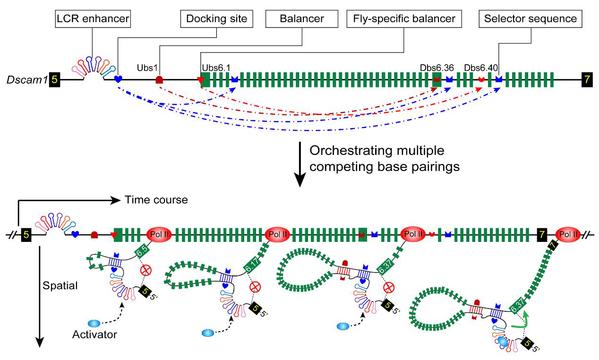
Pancrustacean Dscam1 genes encode 2 000–120 000 distinct isoforms via mutually exclusive splicing; however, the underlying regulatory mechanisms are not fully understood. Here, we revealed a regulatory paradigm of stochastic Dscam1 alternative splicing mediated by conserved long-range RNA structures over 450 million years of evolution. By integrating comparative genomic analysis with mutational evidence, we defined two distinct, evolutionarily conserved RNA structures—the balancer RNA structure and a multi-subunit RNA architecture—that drive the stochastic selection of variable exon 6. The former balances stochastic selection of variable exon 6 isoforms via RNA structural spatial compensation, while the latter acts as a temporal splicing rheostat to counteract the “window of opportunity” for the splicing of 5′-proximal exon 6 variants. Genetic analyses further indicate that these RNA structural elements are essential for proper development and neural wiring. Thus, we proposed a regulatory model in which multiplexed RNA secondary structures coordinate temporally and spatially to orchestrate the stochastic selection of variable exon 6 isoforms. Our findings reveal a new regulatory paradigm of alternative splicing through RNA structural dynamics in cellular processes.
Link: https://doi.org/10.1093/nar/gkag157