Title: A four-dimensional spatial transcriptome atlas of barley caryopsis development and germination
Marta Peirats-Llobet, Zorana Staka, Xiujuan Yang, Yanqiao Zhu, Cunman He, Bhavna Hurgobin, Felipe Ayora, Maria Sofia L Yangzon, Monika W Murcha, Valencia Marisa,
Ghazanfar Abbas Khan, Runxuan Zhang, Iain Milne, Huixia Shou, Matthew R Tucker, Mathew G Lewsey, James Whelan*
Abstract
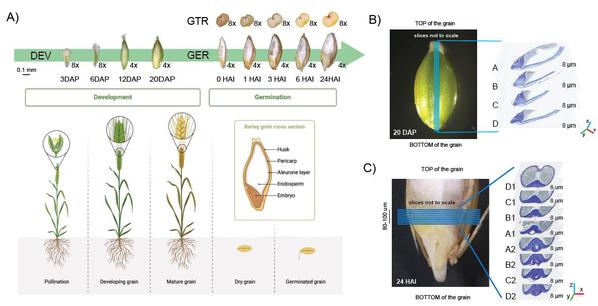
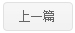
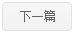
A four-dimensional spatial gene expression atlas of Hordeum vulgare (barley) grain development and germination was generated using spatial transcriptomic analysis of serial sections to reconstruct transcript abundance in three physical dimensions and with temporal kinetics. We investigated the sub-tissue localisations of specific biological activities, using energy biology as an example, including genes encoding proteins involved in starch synthesis and degradation, sugar transport, mitochondrial and chloroplast activity. This atlas revealed different patterns in gene expression across tissues and developmental stages. Heterogeneity in gene expression was observed between clusters, within domains of the individual clusters, across two-dimensional (xy, 55 μm resolution) and three-dimensional (xyz, 8 μm resolution in z-plane) axes. Yet, other genes including typical housekeeping genes such as Actin, Tubulin and others, displayed homogeneous expression patterns. Expression of several genes matched previous gene-specific studies in different barley varieties verifying the robustness of the approach, and indicating that patterns of gene expression are conserved at least for some categories of genes between varieties. Trajectory analysis of aleurone tissue spanning from early development to the completion of germination, provided a comprehensive roadmap of tissue development in terms of processes and identified transcription factors with spatial specificity that play roles in seed development and germination. A public visualization browser is available to view two- and three-dimensional transcription abundance profiles at https://barley4d.latrobe.edu.au/
or https://barley-4d-gene-atlas.hutton.ac.uk/.
Link: https://doi.org/10.1093/plcell/koag120