Title:T2T-Hub: a central platform for analyzing plant and animal telomere-to-telomere genomes
Haoyu chao, Zhonghao Ruan, Zhuojin Li, Xinkai Zhou, Shilong Zhang, Xingxing Shen, Cong Feng, Dijun Chen, Ming Chen
Abstract
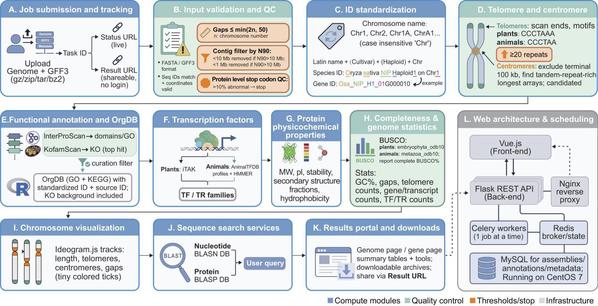
T2T-Hub is a free, web-based platform designed to facilitate the analysis and visualization of telomere-to-telomere (T2T) genomes in plants and animals. The server integrates automated structural and functional annotation workflows with interactive visualization and comparative analysis tools, enabling systematic exploration of complete genome assemblies. T2T-Hub provides a standardized and unified analytical framework specifically tailored for T2T genomes, enabling consistent and comparable analyses across species. To ensure robustness and comparability, we uniformly analyzed 230 high-quality plant and 39 animal T2T genomes using the same standardized workflows, providing users with a consistent reference framework. Users can upload assembled T2T genome sequences together with gene annotation files (GFF3), which are automatically validated and processed through unified analyses including genome quality assessment, telomere and centromere identification, transcription factor prediction, functional annotation, and interactive genome visualization. Importantly, the platform enables interactive genome browsing, searching, and result sharing without requiring any programming skills, thereby lowering the barrier for T2T genome analysis. For internally curated genomes, T2T-Hub additionally provides repeat and noncoding RNA annotation, comparative genomics analyses, and integrated online tools. The platform is freely accessible without login at https://bis.zju.edu.cn/t2thub and https://biobigdata.nju.edu.cn/t2thub.
Link: https://doi.org/10.1093/nar/gkag423